Deposition
The POPSICLE Bacterial Segmentation Dataset: multi-class compartment segmentations for bacterial cryo-ET data
- Deposition ID:CZCDP-10350
Release Date: 2026-05-04
Last Modified: 2026-05-04
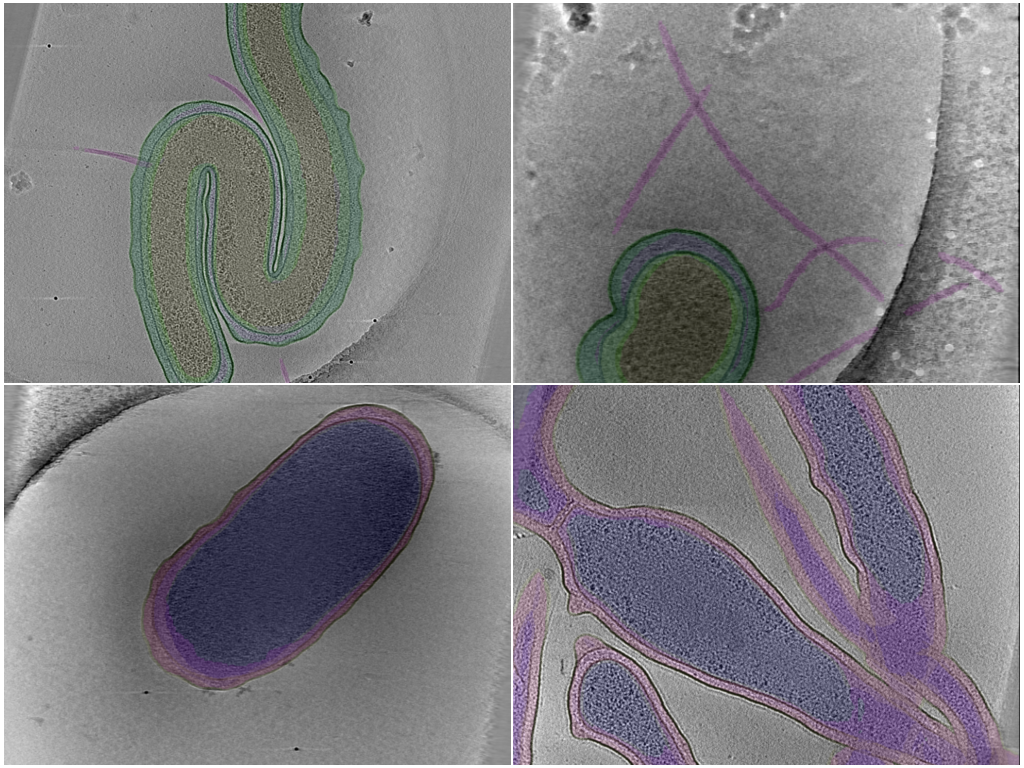
Photo Caption: Central slab of 4 representative tomograms with selected sets of annotations.
Deposition Overview
The bacterial segmentation dataset of POPSICLE (Particle/Object Picking & Segmentation In CryoET Learning & Evaluation), a benchmark suite released to support reproducible evaluation of machine-learning models for dense voxel-wise compartment segmentation in bacterial cryo-ET. The deposition provides semantic segmentations for five classes (cytoplasm, bacterial-type flagellum, membrane, dense body, and periplasmic space / intermembrane space) together with matching 20 Å re-binned WBP tomograms for 80 cryo-ET acquisitions of bacteria drawn from 13 datasets spanning multiple species. Initial annotations were produced semi-manually using napari-nnInteractive, then refined through an agent-designed copick curation pipeline (copick-utils + copick-mcp), followed by manual inspection and correction. Only the final curated segmentation is ingested in this deposition. The accompanying 20 Å tomograms are the resampled volumes used for segmentation (WBP, IMOD reconstruction pipeline; alignment via RAPTOR per the original pipeline) and serve as the visualization base for the segmentations at this voxel spacing.
Authors
- Jonathan Schwartz,
- Utz Heinrich Ermel,
- C. Braxton Owens,
- Zhuowen Zhao,
- Ariana Peck,
- Gus L.W. Hart,
- Grant J. Jensen,
- Bridget Carragher,
- Dari Kimanius
Deposition Data
Annotations:317
Tomograms:80
Publications
Related Databases
Not Submitted
Methods Summary
Annotations | Method Type | Method Details | Method Links |
|---|---|---|---|
| 317 | Hybrid | semi-manual annotation using napari-nnInteractive, followed by an agent-designed copick curation pipeline and manual correction | Source Code:Napari-nnInteractive +2 more |
Deposited Data
Data Contents
Filter by:
317 of 317 Annotations
Annotation Name | Object Shape Type | Method Type | Deposited In |
|---|---|---|---|
104periplasmic space Annotation ID: AN-134577 Ground Truth | SegmentationMask | Hybrid | |
102membrane Annotation ID: AN-134576 Ground Truth | SegmentationMask | Hybrid | |
100cytoplasm Annotation ID: AN-134575 Ground Truth | SegmentationMask | Hybrid | |
104periplasmic space Annotation ID: AN-134574 Ground Truth | SegmentationMask | Hybrid | |
102membrane Annotation ID: AN-134573 Ground Truth | SegmentationMask | Hybrid | |
100cytoplasm Annotation ID: AN-134572 Ground Truth | SegmentationMask | Hybrid | |
104periplasmic space Annotation ID: AN-134571 Ground Truth | SegmentationMask | Hybrid | |
102membrane Annotation ID: AN-134570 Ground Truth | SegmentationMask | Hybrid | |
100cytoplasm Annotation ID: AN-134569 Ground Truth | SegmentationMask | Hybrid | |
104periplasmic space Annotation ID: AN-134568 Ground Truth | SegmentationMask | Hybrid | |
102membrane Annotation ID: AN-134567 Ground Truth | SegmentationMask | Hybrid | |
100cytoplasm Annotation ID: AN-134566 Ground Truth | SegmentationMask | Hybrid | |
104periplasmic space Annotation ID: AN-134565 Ground Truth | SegmentationMask | Hybrid | |
102membrane Annotation ID: AN-134564 Ground Truth | SegmentationMask | Hybrid | |
100cytoplasm Annotation ID: AN-134563 Ground Truth | SegmentationMask | Hybrid | |
104periplasmic space Annotation ID: AN-134562 Ground Truth | SegmentationMask | Hybrid | |
102membrane Annotation ID: AN-134561 Ground Truth | SegmentationMask | Hybrid | |
100cytoplasm Annotation ID: AN-134560 Ground Truth | SegmentationMask | Hybrid | |
104periplasmic space Annotation ID: AN-134559 Ground Truth | SegmentationMask | Hybrid | |
102membrane Annotation ID: AN-134558 Ground Truth | SegmentationMask | Hybrid |
- 1
- 2
- 3
- 4
- 5
- 32